Note
Go to the end to download the full example code.
Gradient Flow Field Visualization¶
This example creates a mesmerizing flow field visualization with gradient colors that demonstrates dartwork-mpl’s advanced color interpolation and styling capabilities.
import matplotlib.patheffects as path_effects
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.colors import LinearSegmentedColormap
import dartwork_mpl as dm
# Set random seed for reproducibility
np.random.seed(42)
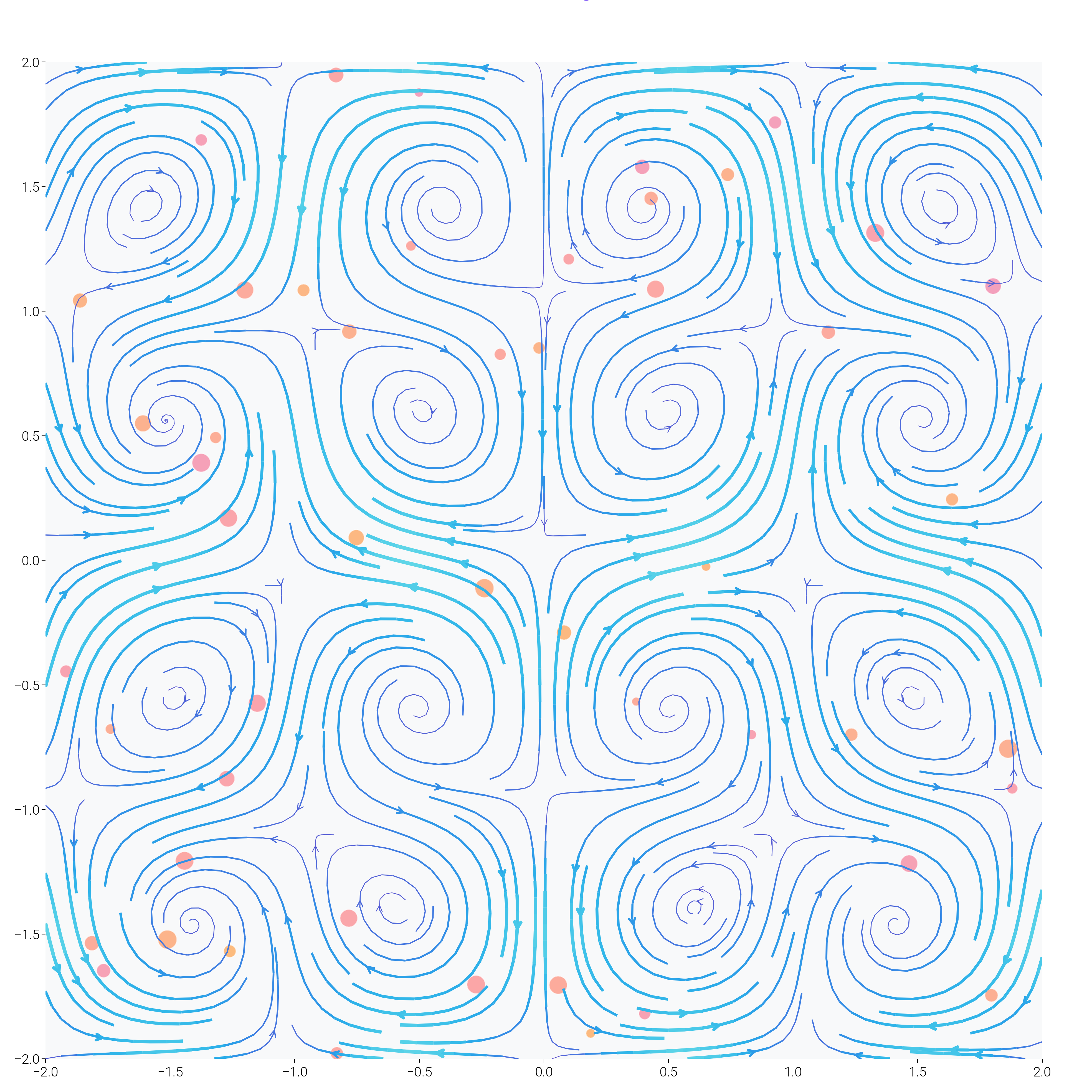
Organic Flow Field with Color Gradients¶
Create a flow field that looks like organic patterns with smooth color transitions.
dm.style.use("scientific")
# Create a grid of vectors
fig, ax = plt.subplots(figsize=(dm.cm2in(20), dm.cm2in(20)))
# Generate flow field
n_points = 25
x = np.linspace(-2, 2, n_points)
y = np.linspace(-2, 2, n_points)
X, Y = np.meshgrid(x, y)
# Create interesting flow patterns
t = 2.5
U = np.sin(np.pi * X) * np.cos(np.pi * Y) + 0.3 * np.sin(t * X)
V = -np.cos(np.pi * X) * np.sin(np.pi * Y) + 0.3 * np.cos(t * Y)
# Calculate magnitude for colors
M = np.sqrt(U**2 + V**2)
# Create custom colormap using dartwork's color interpolation
colors_flow = dm.cspace("oc.violet9", "oc.cyan3", n=256, space="oklch")
flow_cmap = LinearSegmentedColormap.from_list(
"flow", [c.to_hex() for c in colors_flow]
)
# Plot streamlines with varying thickness
strm = ax.streamplot(
X,
Y,
U,
V,
color=M,
cmap=flow_cmap,
linewidth=2 * M / M.max(),
density=2,
arrowsize=0.8,
arrowstyle="->",
)
# Add particles flowing along the field
n_particles = 50
px = np.random.uniform(-2, 2, n_particles)
py = np.random.uniform(-2, 2, n_particles)
particle_colors = dm.cspace("oc.pink5", "oc.orange5", n=n_particles)
for _i, (x_p, y_p, color) in enumerate(zip(px, py, particle_colors, strict=False)):
size = np.random.uniform(20, 100)
ax.scatter(
x_p,
y_p,
s=size,
c=[color.to_hex()],
alpha=0.6,
edgecolors="white",
linewidths=0.5,
)
# Style the plot
dm.hide_all_spines(ax)
ax.set_xlim(-2, 2)
ax.set_ylim(-2, 2)
ax.set_aspect("equal")
ax.set_facecolor("oc.gray0")
# Add title with glow effect
title = ax.text(
0,
2.3,
"Quantum Flow Dynamics",
ha="center",
va="center",
fontsize=dm.fs(4),
fontweight=dm.fw(3),
color="white",
)
title.set_path_effects(
[path_effects.withStroke(linewidth=3, foreground="oc.violet5")]
)
dm.simple_layout(fig)

Radial Burst Pattern with Layered Gradients¶
Create a radial visualization that combines multiple gradient layers.
fig, ax = plt.subplots(
figsize=(dm.cm2in(18), dm.cm2in(18)), subplot_kw={"projection": "polar"}
)
# Generate radial data
theta = np.linspace(0, 2 * np.pi, 360)
r_layers = 8
for layer in range(r_layers):
# Create varying radius for each layer
r_base = 0.5 + layer * 0.5
r = r_base + 0.3 * np.sin(6 * theta + layer * np.pi / 4)
# Color gradient for each layer
layer_colors = dm.cspace(
f"oc.{['blue', 'teal', 'green', 'yellow', 'orange', 'red', 'pink', 'violet'][layer]}5",
f"oc.{['blue', 'teal', 'green', 'yellow', 'orange', 'red', 'pink', 'violet'][layer]}8",
n=len(theta),
)
# Plot with transparency
for i in range(len(theta) - 1):
ax.fill_between(
[theta[i], theta[i + 1]],
0,
[r[i], r[i + 1]],
color=layer_colors[i].to_hex(),
alpha=0.3 + 0.05 * layer,
)
# Add radial spokes
for angle in np.linspace(0, 2 * np.pi, 12, endpoint=False):
ax.plot([angle, angle], [0, 4], color="white", alpha=0.2, lw=0.5)
# Styling
ax.set_ylim(0, 4)
ax.set_theta_zero_location("N")
ax.set_theta_direction(-1)
dm.hide_all_spines(ax)
ax.set_xticks([])
ax.set_yticks([])
ax.set_facecolor("oc.gray9")
# Add center burst
center = plt.Circle(
(0, 0), 0.3, transform=ax.transData._b, color="white", zorder=10
)
ax.add_patch(center)
ax.text(
0,
0,
"✦",
fontsize=dm.fs(6),
ha="center",
va="center",
color="oc.gray9",
weight="bold",
zorder=11,
)
dm.simple_layout(fig)
DNA Helix Visualization with Gradient Ribbons¶
Create a 3D-like DNA helix using creative 2D techniques.
fig, ax = plt.subplots(figsize=(dm.cm2in(12), dm.cm2in(20)))
# Generate helix coordinates
t = np.linspace(0, 4 * np.pi, 500)
x1 = np.sin(t)
x2 = np.sin(t + np.pi)
y = t
# Create gradient colors along the helix
colors_helix1 = dm.cspace("oc.blue3", "oc.purple7", n=len(t))
colors_helix2 = dm.cspace("oc.green3", "oc.orange7", n=len(t))
# Draw the helixes with varying width to create depth
for i in range(len(t) - 1):
# First strand
width1 = 2 + np.sin(t[i]) * 0.5 # Varying width for 3D effect
ax.plot(
[x1[i], x1[i + 1]],
[y[i], y[i + 1]],
color=colors_helix1[i].to_hex(),
linewidth=width1,
alpha=0.8,
solid_capstyle="round",
)
# Second strand
width2 = 2 - np.sin(t[i]) * 0.5
ax.plot(
[x2[i], x2[i + 1]],
[y[i], y[i + 1]],
color=colors_helix2[i].to_hex(),
linewidth=width2,
alpha=0.8,
solid_capstyle="round",
)
# Connect strands at intervals
if i % 25 == 0:
ax.plot(
[x1[i], x2[i]],
[y[i], y[i]],
color="white",
alpha=0.3,
lw=0.5,
linestyle=":",
)
# Add base pair representations
for i in range(0, len(t), 40):
# Adenine-Thymine pairs
if i % 80 == 0:
ax.scatter(
[x1[i], x2[i]],
[y[i], y[i]],
c=["oc.red5", "oc.blue5"],
s=100,
edgecolors="white",
linewidths=1,
zorder=5,
)
# Guanine-Cytosine pairs
else:
ax.scatter(
[x1[i], x2[i]],
[y[i], y[i]],
c=["oc.green5", "oc.yellow5"],
s=100,
edgecolors="white",
linewidths=1,
zorder=5,
)
# Styling
dm.hide_all_spines(ax)
ax.set_xlim(-2, 2)
ax.set_ylim(-0.5, 4 * np.pi + 0.5)
ax.set_aspect("equal")
ax.set_facecolor("oc.gray1")
# Add labels
ax.text(
0,
-1,
"DNA Double Helix",
ha="center",
fontsize=dm.fs(3),
color="white",
weight="bold",
)
ax.text(
0,
4 * np.pi + 1,
"A-T G-C Base Pairs",
ha="center",
fontsize=dm.fs(0),
color="oc.gray5",
)
dm.simple_layout(fig)
Fibonacci Spiral Galaxy¶
Create a galaxy visualization based on the Fibonacci spiral.
fig, ax = plt.subplots(figsize=(dm.cm2in(18), dm.cm2in(18)))
# Generate Fibonacci spiral
def fibonacci_spiral(n_points=1000):
golden_ratio = (1 + np.sqrt(5)) / 2
theta = np.linspace(0, 8 * np.pi, n_points)
r = np.exp(theta / (2 * np.pi / np.log(golden_ratio)))
x = r * np.cos(theta)
y = r * np.sin(theta)
return x, y, theta
x, y, theta = fibonacci_spiral(1500)
# Normalize for colors
norm_theta = (theta - theta.min()) / (theta.max() - theta.min())
# Create spiral galaxy effect with multiple arms
for arm in range(3):
angle_offset = arm * 2 * np.pi / 3
x_arm = x * np.cos(angle_offset) - y * np.sin(angle_offset)
y_arm = x * np.sin(angle_offset) + y * np.cos(angle_offset)
# Color gradient for each arm
arm_colors = dm.cspace(
f"oc.{['indigo', 'violet', 'blue'][arm]}9",
f"oc.{['cyan', 'teal', 'green'][arm]}3",
n=len(x),
)
# Draw arm with varying size points
for i in range(0, len(x), 2):
size = 0.5 + 20 * np.exp(-theta[i] / 10)
alpha = 0.8 * np.exp(-theta[i] / 15)
ax.scatter(
x_arm[i],
y_arm[i],
s=size,
c=[arm_colors[i].to_hex()],
alpha=alpha,
edgecolors="none",
)
# Add star clusters
n_stars = 500
star_theta = np.random.uniform(0, 8 * np.pi, n_stars)
star_r = np.exp(star_theta / (2 * np.pi / np.log((1 + np.sqrt(5)) / 2)))
star_r *= np.random.uniform(0.8, 1.2, n_stars) # Add randomness
star_x = star_r * np.cos(star_theta)
star_y = star_r * np.sin(star_theta)
ax.scatter(
star_x,
star_y,
s=np.random.uniform(0.1, 2, n_stars),
c="white",
alpha=np.random.uniform(0.3, 1, n_stars),
)
# Add galaxy center
center_glow = plt.Circle((0, 0), 30, color="white", alpha=0.05)
ax.add_patch(center_glow)
center_glow2 = plt.Circle((0, 0), 20, color="oc.yellow3", alpha=0.1)
ax.add_patch(center_glow2)
center_core = plt.Circle((0, 0), 10, color="oc.yellow5", alpha=0.3)
ax.add_patch(center_core)
# Styling
dm.hide_all_spines(ax)
ax.set_xlim(-150, 150)
ax.set_ylim(-150, 150)
ax.set_aspect("equal")
ax.set_facecolor("black")
# Add title
title_text = ax.text(
0,
-170,
"Fibonacci Galaxy M-" + str(np.random.randint(100, 999)),
ha="center",
fontsize=dm.fs(3),
color="white",
alpha=0.8,
)
title_text.set_path_effects(
[path_effects.withStroke(linewidth=2, foreground="oc.blue9")]
)
dm.simple_layout(fig)
Generative Art: Voronoi Tessellation with Gradients¶
Create artistic Voronoi patterns with smooth color transitions.
from matplotlib.patches import Polygon # noqa: E402
from scipy.spatial import Voronoi # noqa: E402
fig, ax = plt.subplots(figsize=(dm.cm2in(20), dm.cm2in(20)))
# Generate random points with clustering
n_clusters = 5
n_points_per_cluster = 8
points = []
for _ in range(n_clusters):
center_x = np.random.uniform(-8, 8)
center_y = np.random.uniform(-8, 8)
cluster_points = np.random.randn(n_points_per_cluster, 2) * 1.5 + [
center_x,
center_y,
]
points.extend(cluster_points)
points = np.array(points)
# Add boundary points to ensure closed regions
boundary_points = []
for x in np.linspace(-10, 10, 10):
boundary_points.append([x, -10])
boundary_points.append([x, 10])
for y in np.linspace(-10, 10, 10):
boundary_points.append([-10, y])
boundary_points.append([10, y])
all_points = np.vstack([points, boundary_points])
# Compute Voronoi tessellation
vor = Voronoi(all_points)
# Color each region with gradient
for i, region in enumerate(vor.regions):
if not region or -1 in region:
continue
polygon = [vor.vertices[j] for j in region]
if len(polygon) > 2:
# Check if polygon is within bounds
polygon_array = np.array(polygon)
if (
polygon_array[:, 0].min() < -10
or polygon_array[:, 0].max() > 10
or polygon_array[:, 1].min() < -10
or polygon_array[:, 1].max() > 10
):
continue
# Create color based on position and index
center = polygon_array.mean(axis=0)
hue = (np.arctan2(center[1], center[0]) + np.pi) / (2 * np.pi) * 360
color = dm.oklch(0.6 + 0.2 * np.sin(i), 0.2, hue)
poly = Polygon(
polygon,
facecolor=color.to_hex(),
edgecolor="white",
linewidth=0.5,
alpha=0.8,
)
ax.add_patch(poly)
# Add decorative elements
for point in points[: n_clusters * n_points_per_cluster]:
ax.scatter(
*point, s=20, c="white", edgecolors="oc.gray7", linewidths=1, zorder=10
)
# Styling
ax.set_xlim(-10, 10)
ax.set_ylim(-10, 10)
ax.set_aspect("equal")
dm.hide_all_spines(ax)
ax.set_facecolor("oc.gray1")
# Title
ax.text(
0,
-11,
"Voronoi Dreams",
ha="center",
fontsize=dm.fs(3),
color="white",
weight="bold",
alpha=0.9,
)
dm.simple_layout(fig)
plt.show()
Total running time of the script: (0 minutes 42.141 seconds)