Note
Go to the end to download the full example code.
Bayesian MCMC Posterior Ridgelines¶
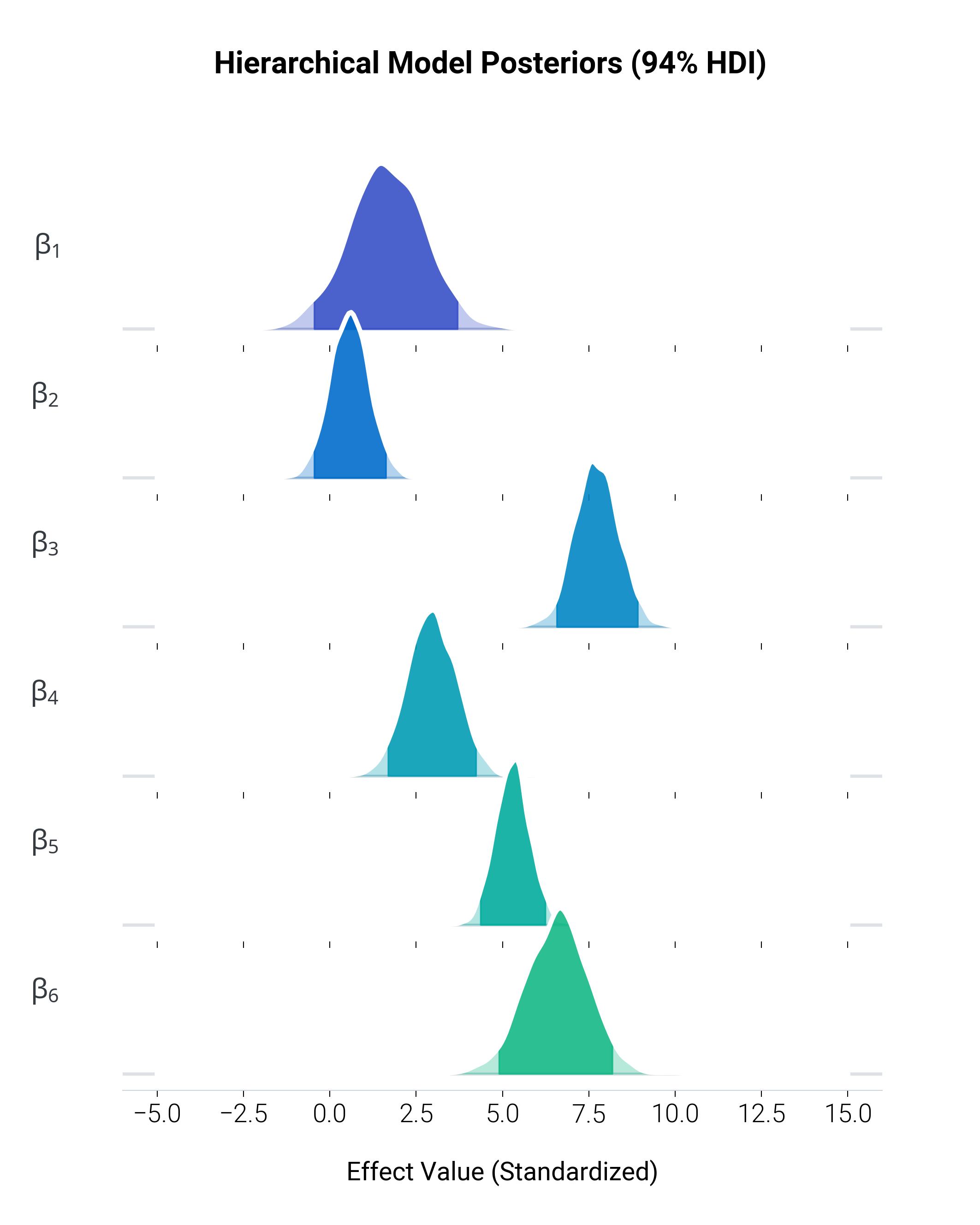
Visualizing the posterior distributions of multiple parameters sampled via Markov Chain Monte Carlo (MCMC). This “joyplot” or ridgeline plot effectively compares parameter uncertainties.
In Bayesian workflows, it’s customary to highlight the Highest Density Interval (HDI).
Here, we overlay a global cspace gradient across the parameters, shading the
bulk of the distributions to give depth and immediate visual separation.
import matplotlib.pyplot as plt
import numpy as np
from scipy.stats import gaussian_kde
import dartwork_mpl as dm
dm.style.use("report")
# Generate synthetic MCMC trace data for 6 parameters
np.random.seed(101)
parameters = [f"$\\beta_{i}$" for i in range(1, 7)]
n_params = len(parameters)
traces = []
for i in range(n_params):
mean = np.random.uniform(-2, 5) + (i * 1.5)
std = np.random.uniform(0.5, 1.5)
traces.append(np.random.normal(mean, std, 3000))
# Colors extracted from an OKLCH perceptual space
c1 = dm.named("oc.indigo9").to_hex()
c2 = dm.named("oc.teal6").to_hex()
ridge_colors = dm.cspace(c1, c2, n=n_params, space="oklch")
fig, axes = plt.subplots(
n_params, 1, figsize=(dm.SW * 1.2, dm.SW * 1.5), sharex=True
)
fig.subplots_adjust(
hspace=-0.25
) # Negative hspace for the overlapping ridgeline effect
x_eval = np.linspace(-5, 15, 300)
for _i, (ax, trace, color, label) in enumerate(
zip(axes, traces, ridge_colors, parameters, strict=False)
):
kde = gaussian_kde(trace)
density = kde(x_eval)
# Scale density for uniform visual height in the overlap
density_scaled = density / density.max()
# Extract the 94% HDI
lo = np.percentile(trace, 3)
hi = np.percentile(trace, 97)
# White outline
ax.plot(x_eval, density_scaled, color="white", lw=1.5, zorder=5)
# Fill the entire distribution lightly
ax.fill_between(
x_eval, 0, density_scaled, color=color.to_hex(), alpha=0.3, zorder=4
)
# Fill the HDI bounds heavily
hdi_mask = (x_eval >= lo) & (x_eval <= hi)
ax.fill_between(
x_eval,
0,
density_scaled,
where=hdi_mask,
color=color.to_hex(),
alpha=0.85,
zorder=4,
)
# Cleanup individual axes
for spine in ax.spines.values():
spine.set_visible(False)
ax.set_yticks([])
ax.set_ylabel(
label,
rotation=0,
labelpad=20,
ha="right",
va="center",
fontsize=dm.fs(0.5),
weight="bold",
color="oc.gray8",
)
ax.patch.set_alpha(0) # Transparent background
ax.axhline(0, color="oc.gray3", lw=1, zorder=3)
# Shared x-axis on the bottom
axes[-1].spines["bottom"].set_visible(True)
axes[-1].spines["bottom"].set_color("oc.gray4")
axes[-1].set_xlabel(
"Effect Value (Standardized)", fontsize=dm.fs(0), labelpad=10
)
fig.suptitle(
"Hierarchical Model Posteriors (94% HDI)",
fontsize=dm.fs(1.5),
weight="bold",
y=0.96,
)
Total running time of the script: (0 minutes 1.525 seconds)